Projects
We offer a variety of exciting projects for students, ranging from investigating the molecular mechanisms of protein synthesis in chloroplasts to developing innovative tools for synthetic biology. Our projects provide hands-on experience with cutting-edge techniques and the opportunity to contribute to impactful research in plant biology.
Our projects
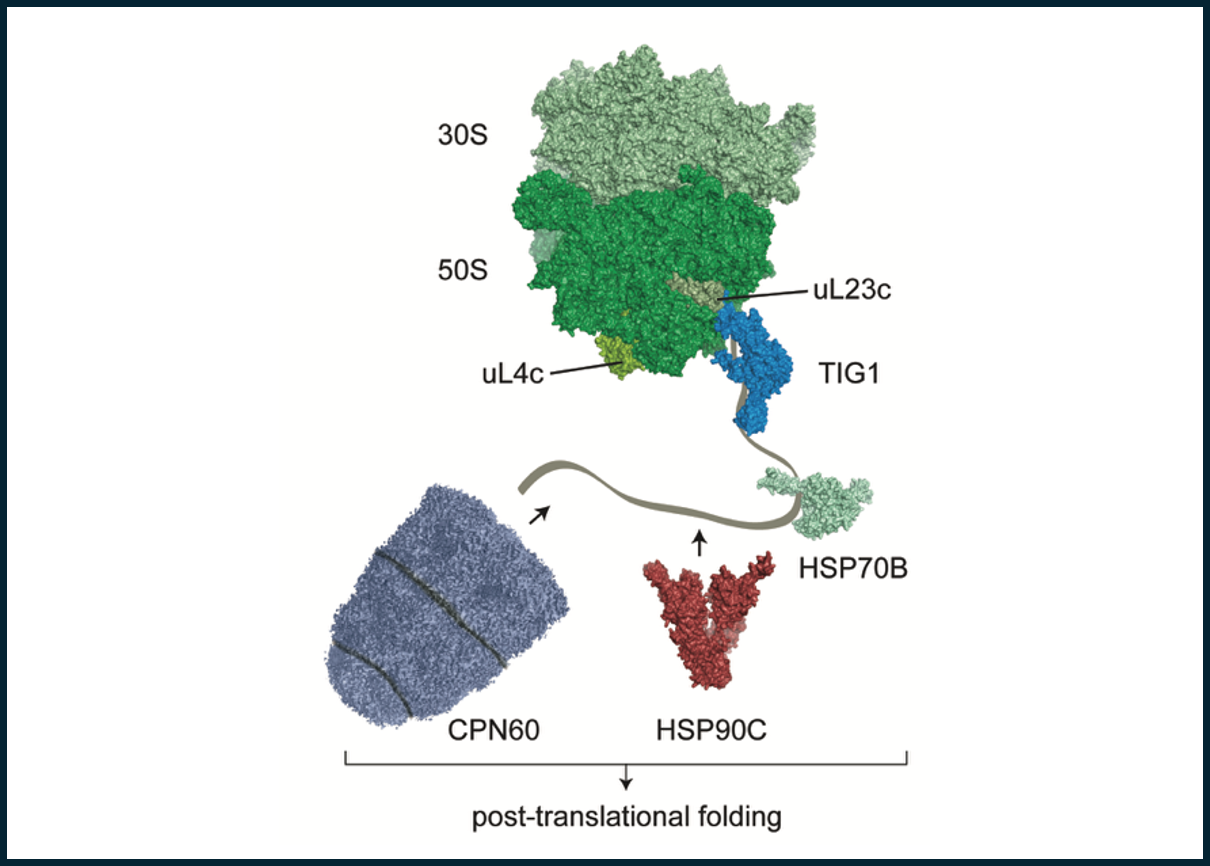
Throughout their lifespan, proteins are guided by a number of factors that facilitate their maturation and that help proteins to gain their functionality even under stress and denaturing conditions. Many of these factors belong to the family of molecular chaperones. We are currently focusing on two molecular chaperones that act early during maturation of a nascent polypeptide. We investigated a previously overlooked protein, termed trigger factor. Trigger factor has a dragon-shaped conformation and binds on translating ribosomes, presumable to assist the emerging polypeptide to achieve folding. We are currently trying to understand which of the essential chloroplast-encoded subunits of the photosynthesis machinery require this chaperone during maturation. In addition, we try to understand the mechanism of the CPN60 machinery. CPN60 contains two stacked rings of 7 subunits each and it is known that this chaperon is important for Rubisco folding. Our goal is to understand, why CPN60 evolved a much more complicated composition when compared to its bacterial ortholog and how this contributes to the folding requirements within the chloroplast.
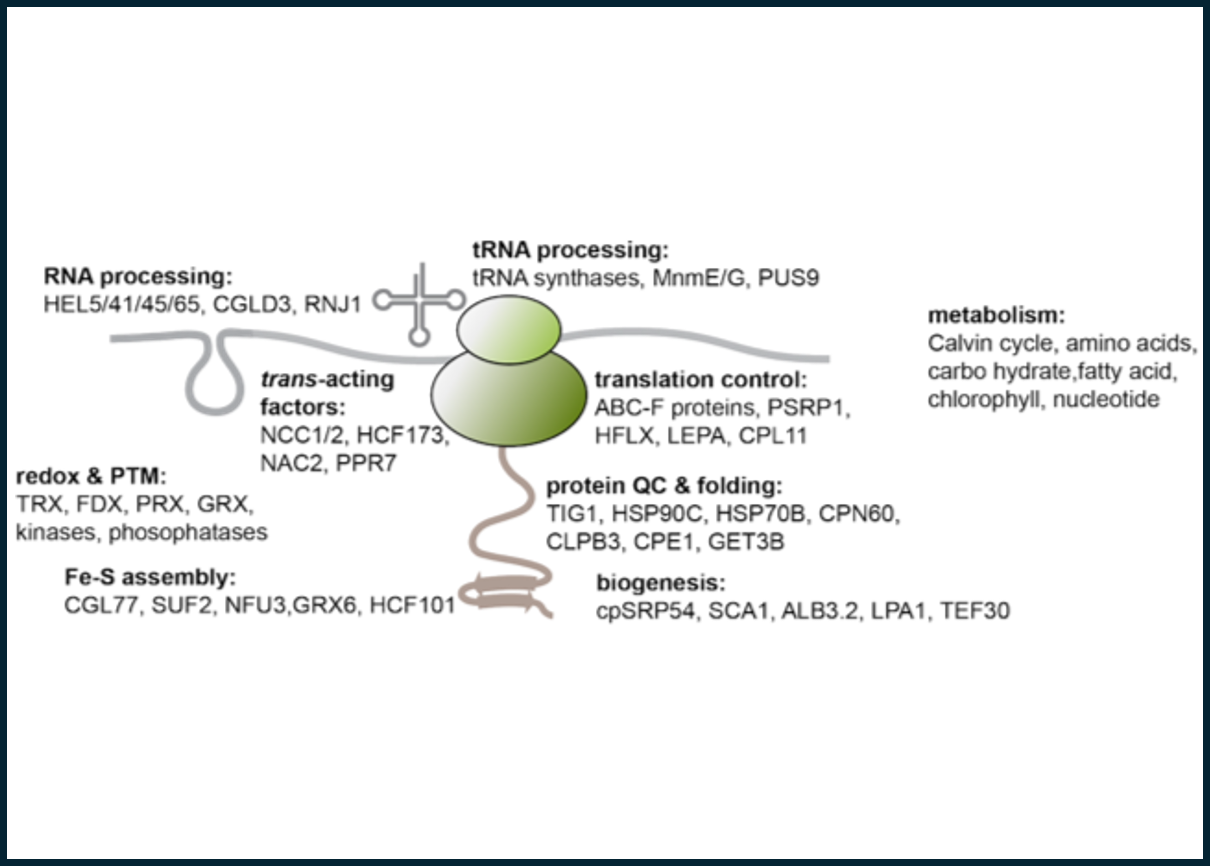
After endosymbiosis, many genes of the former cyanobacteria-like endosymbiont were transferred from the chloroplast to the nucleus of the plant cell. However, several genes remained on the plastidic genome, mainly comprising essential subunits of the photosynthesis machinery and the gene expression apparatus. Interestingly, expression of these genes is strictly regulated on a co-translational level, presumably for allowing quick and efficient adjustments to compensate environmental changes. Previously, we aimed to understand which proteins interact with the chloroplast ribosome, in order to better understand which players may conduct a regulatory function. We identified a large network of proteins and could show how the chloroplast ribosome is closely embedded into several biochemical pathways. We are now aiming to better understand the function of several of these identified factors. Moreover, the chloroplast protein synthesis machinery is an intriguing process for quantitative modelling due to its reduction (e.g. lower number of tRNAs), when compared to other systems. We explore the underlying rates of translation that will allow to comprehensively determine dynamics and limitations of protein synthesis.
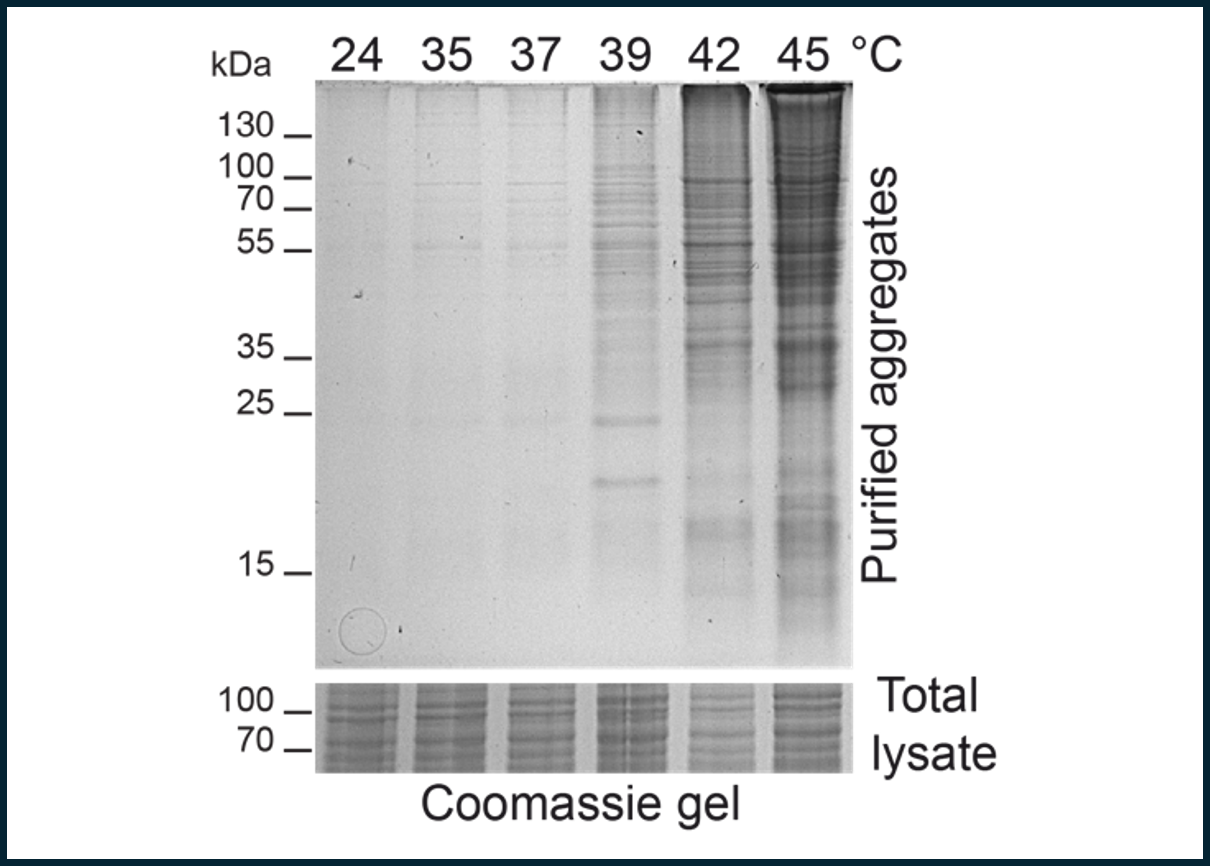
Temperature stress such as sudden heat exposure has drastic consequences for protein homeostasis (protein folding, solubility, activity and half-life). By challenging protein homeostasis, proteins are prone to misfolding and aggregation, with deleterious consequences for the overall cellular physiology. We observed that heat exposure leads to wide-spread aggregation in plant cells. We are now aiming to understand why proteins aggregate and which biochemical pathways are most affected by heat-induced aggregation. Furthermore, we try to understand regulatory processes in plant cells (mainly in the chloroplast) that help to fast and efficient acclimation to changed environmental conditions. By this we aim to reveal central player of acclimation, which may help to engineer plants towards a higher stress resilience.
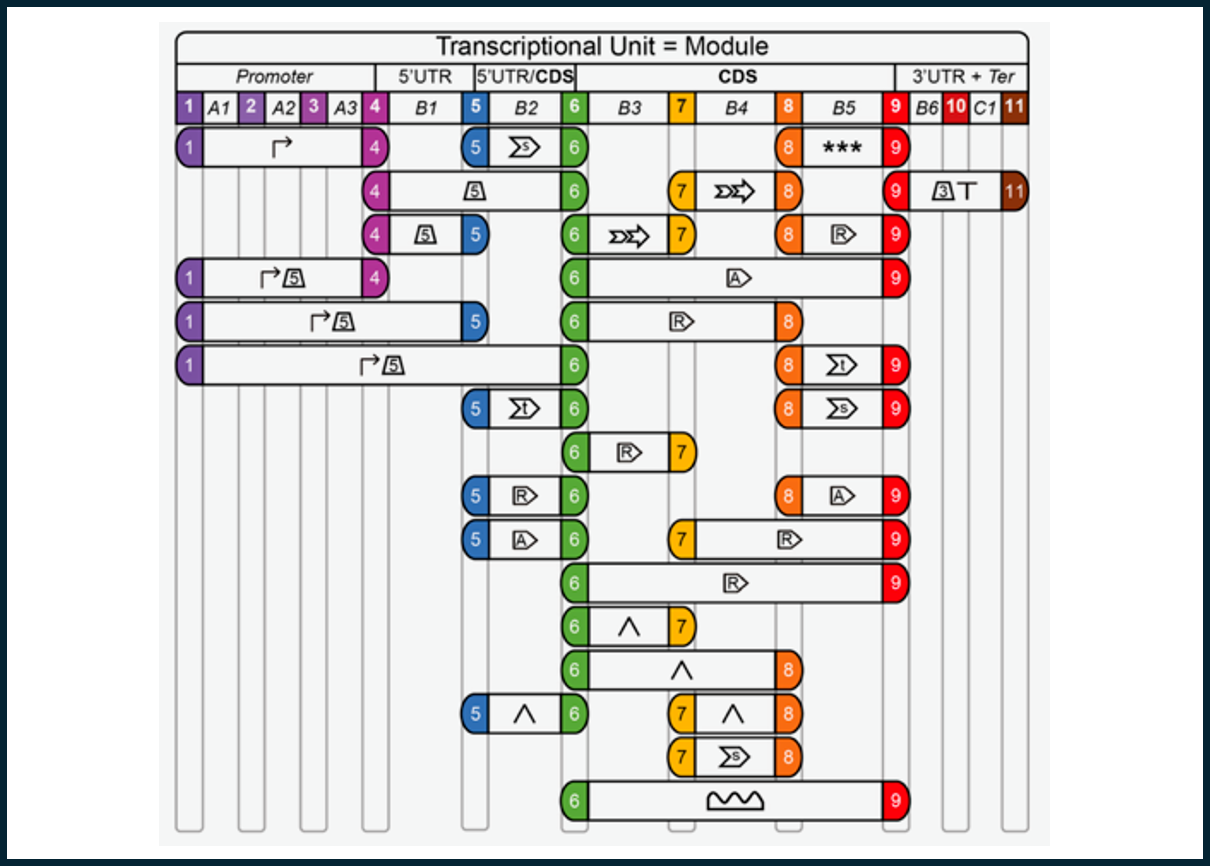
Over the past years, we have been collaborating with several other laboratories, word-wide, in order to establish robust tools for synthetic biology with the unicellular green alga Chlamydomonas reinhardtii. For expression of transgenes from the nuclear genome, we contributed to collect a large set of genetic elements, selection markers, reporters and affinity tags, which can be efficiently assembled into transcriptional units via the standardized Modular Cloning (MoClo) procedure. This greatly improved transgene expression in the alga and allows to quickly test various constructs for optimal transgene expression. Moreover, we contribute to setting up a MoClo-based platform for engineering of the chloroplast genome. Thus, we have all tools in hand that allows us to explore synthetic biology of the nuclear and the chloroplast genome of C. reinhardtii. In addition, we employ Crispr/Cas for engineering of the nuclear genome.